Abstract
An Arabidopsis mutant named drol1 was originally isolated as a mutant with defects in the repression of OLEOSIN1 after seed germination. DROL1 encodes a homologous protein to DIB1, a subunit of U5 snRNP in the spliceosome. Comprehensive analysis of intron splicing by RNA-Seq analysis of drol1 mutants revealed reduced splicing of most of the minor introns with AT–AC dinucleotide termini. This suggested that the phenotype of drol1 was due to insufficient expression of genes with AT–AC introns.
We screened for secondary mutations which suppress the drol1 morphological phenotypes, and seven suppressors were isolated, all of which had mutations in U5 snRNP subunits. Although the morphology of the suppressors was almost similar to wild type, the extent of splicing recovery varied. Detailed analysis revealed that AT–AC introns can be spliced by spliceosomes that normally process GT–AG introns. Furthermore, mRNAs retaining introns were found to be retained in the nucleus. These findings indicate that the drol1 phenotypes are not solely due to dysfunction of genes containing AT–AC introns, but also involve ABA-like signaling triggered by the accumulation of intron-retaining mRNAs.
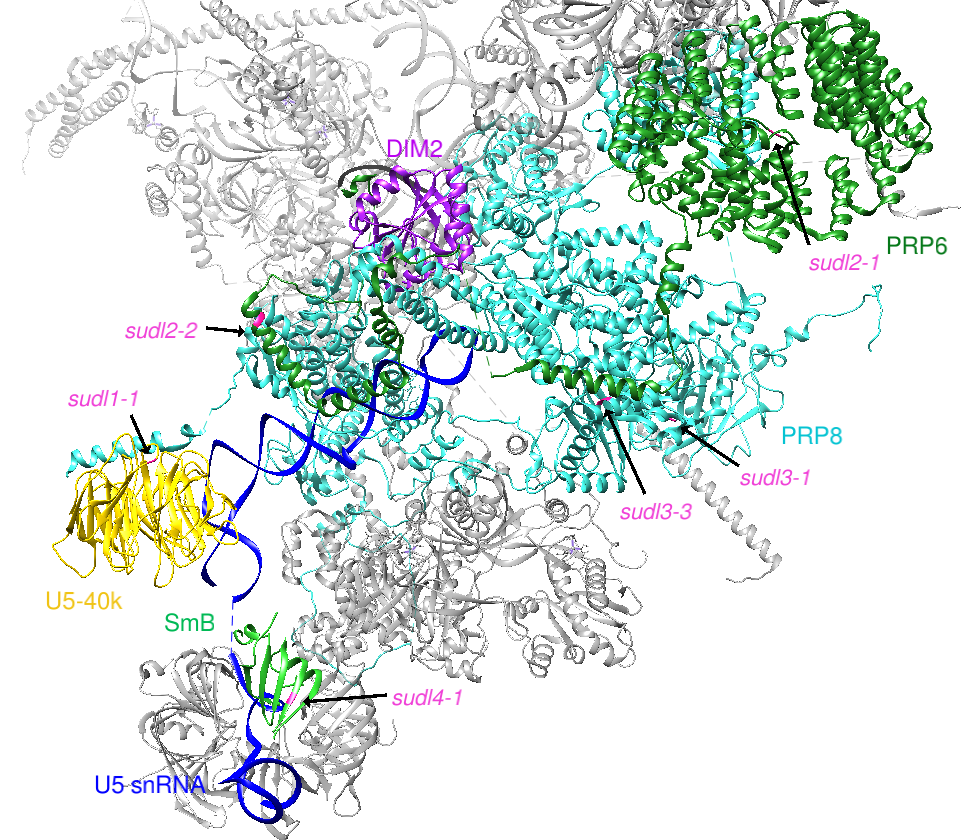
Background: DROL1 Encodes a Splicing Factor in U5 snRNP, Which Is Specifically Required for AT–AC-type Introns
DROL1 and DIB1 Form Distinct Subfamilies
Multiple alignment
Phylogenetic tree
- Arabidopsis DROL1 is a homolog of yeast DIB1 but belongs to a different subfamily.
- All eukaryotes have more than one homolog of DIB1, but DROL1 is restricted to a part of eukaryotes.
Specific Retention of AT–AC Introns in drol1
WT
drol1-1
- RNA-Seq reads were aligned to HD2B, whose third intron is AT–AC.
- In the drol1-1 mutant, this third AT–AC intron is specifically retained.
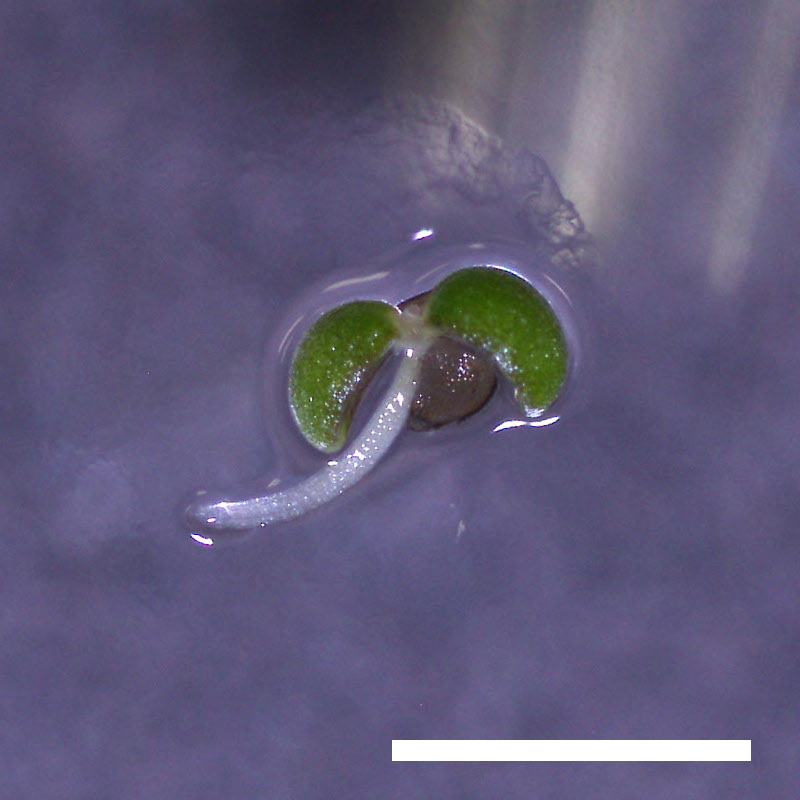
drol1 Is Hypersensitive to ABA
| ABA | |||||
|---|---|---|---|---|---|
| 0 µM | 0.5 µM | 1 µM | 2 µM | 5 µM | |
| Col |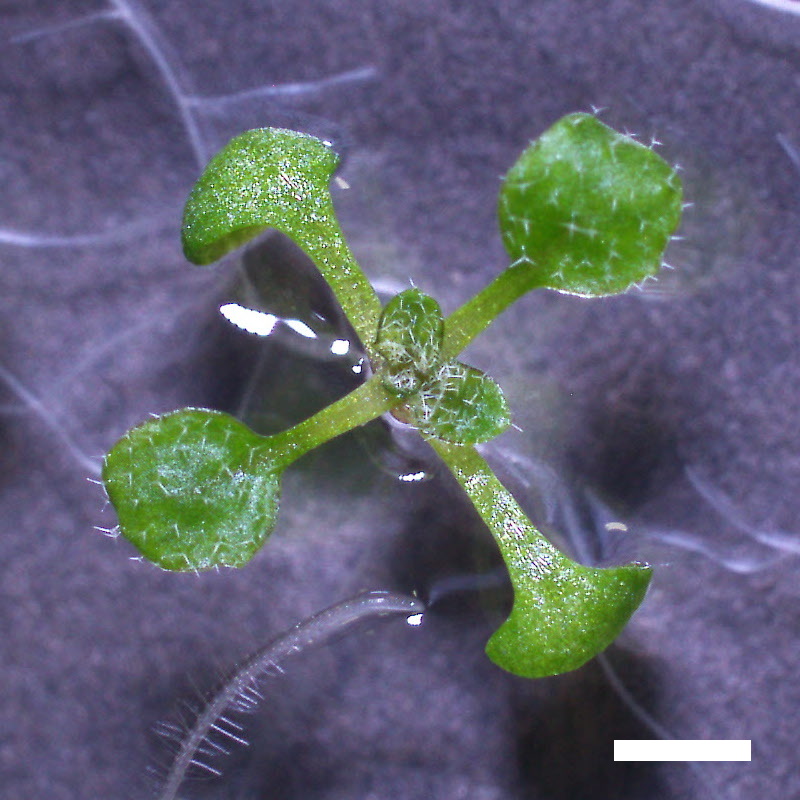  |
 |
 |
 |
 |
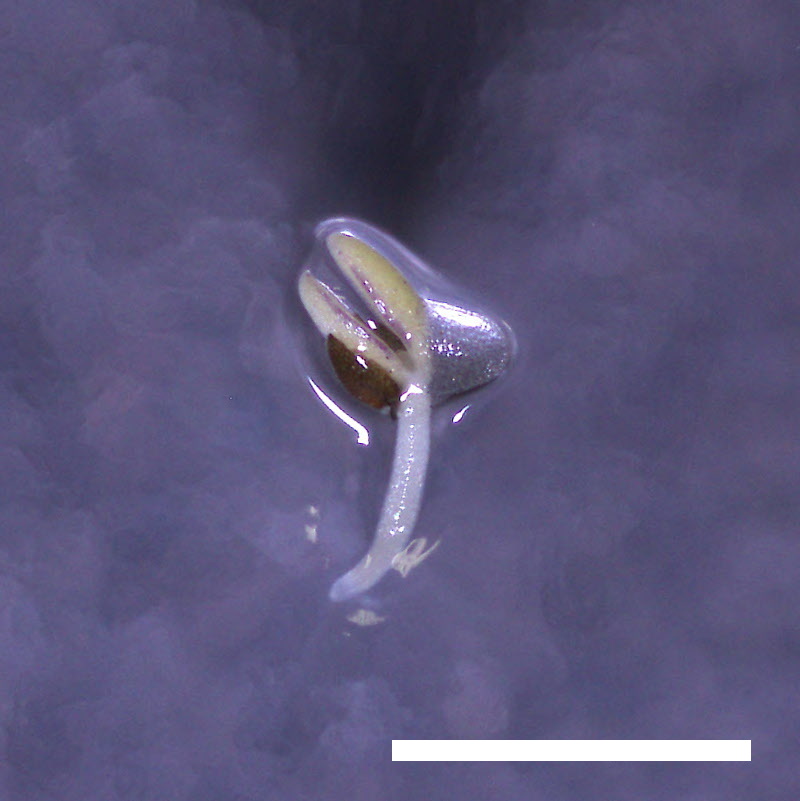
| drol1-1 |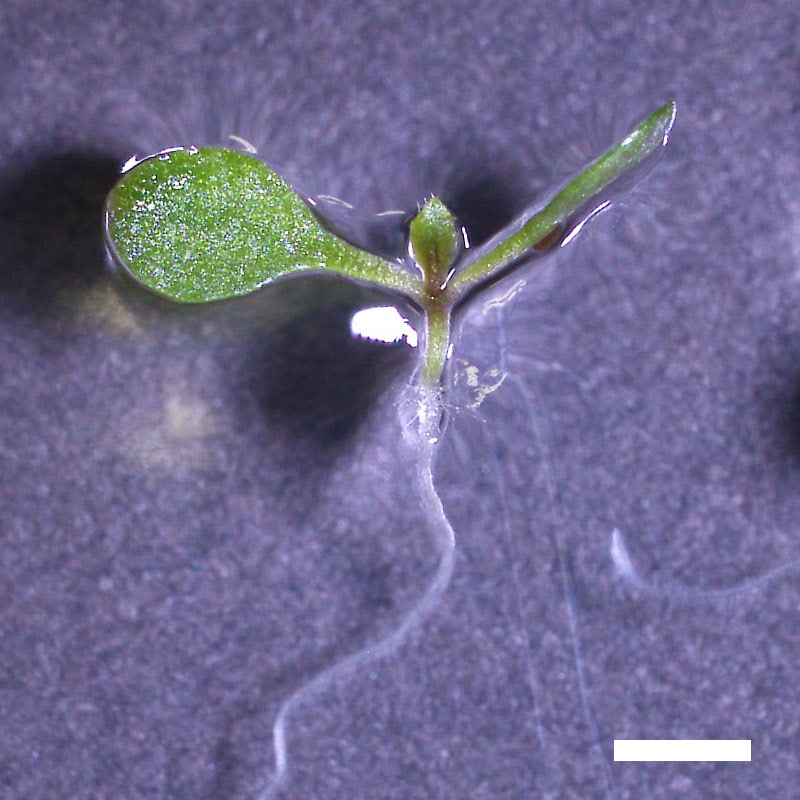  |
 |
 |
 |
 |
| drol1-2 |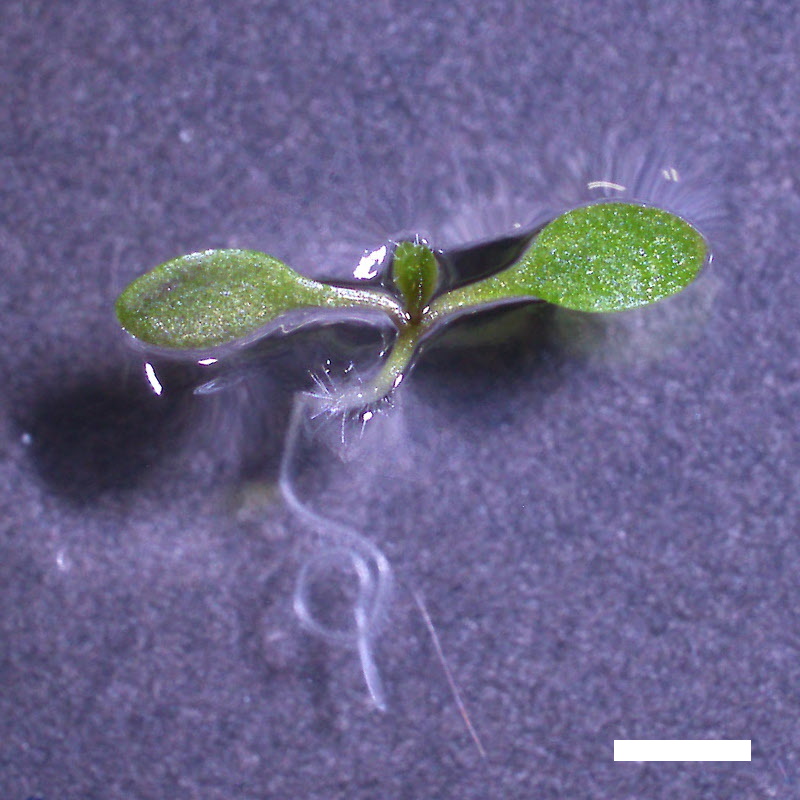  |
 |
 |
 |
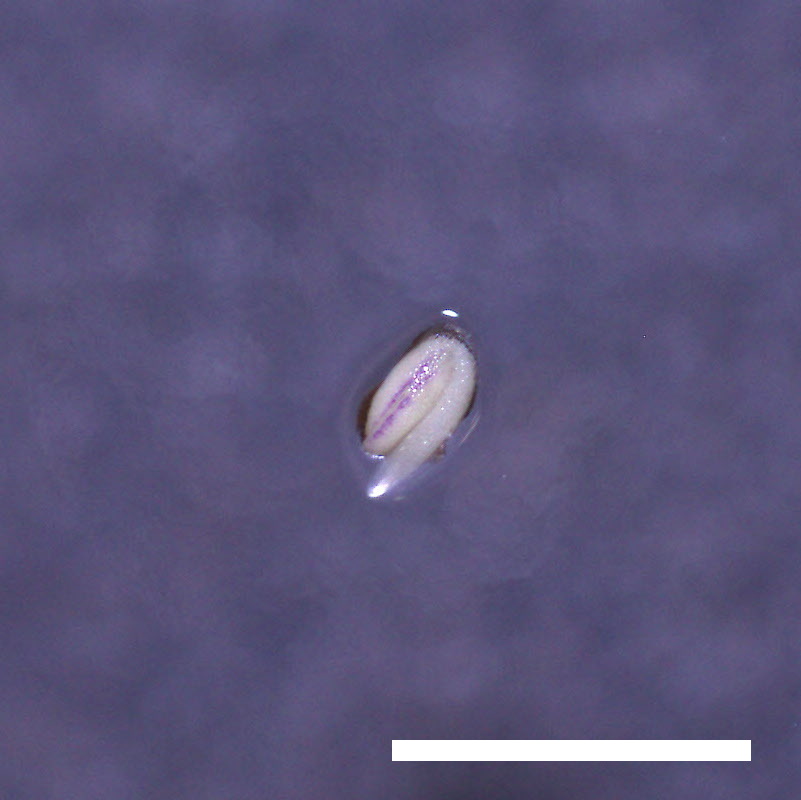 |
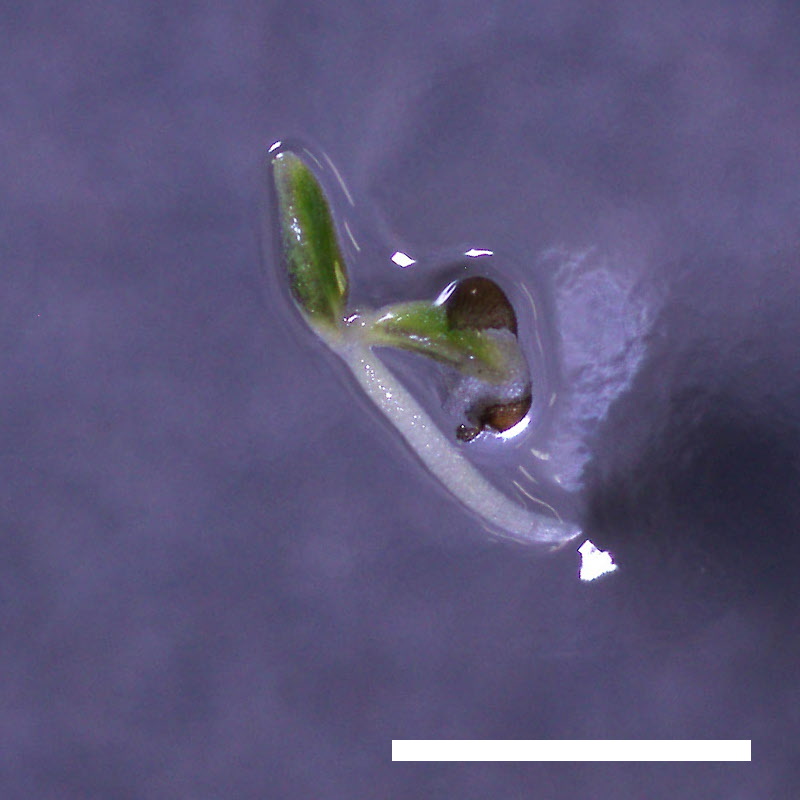
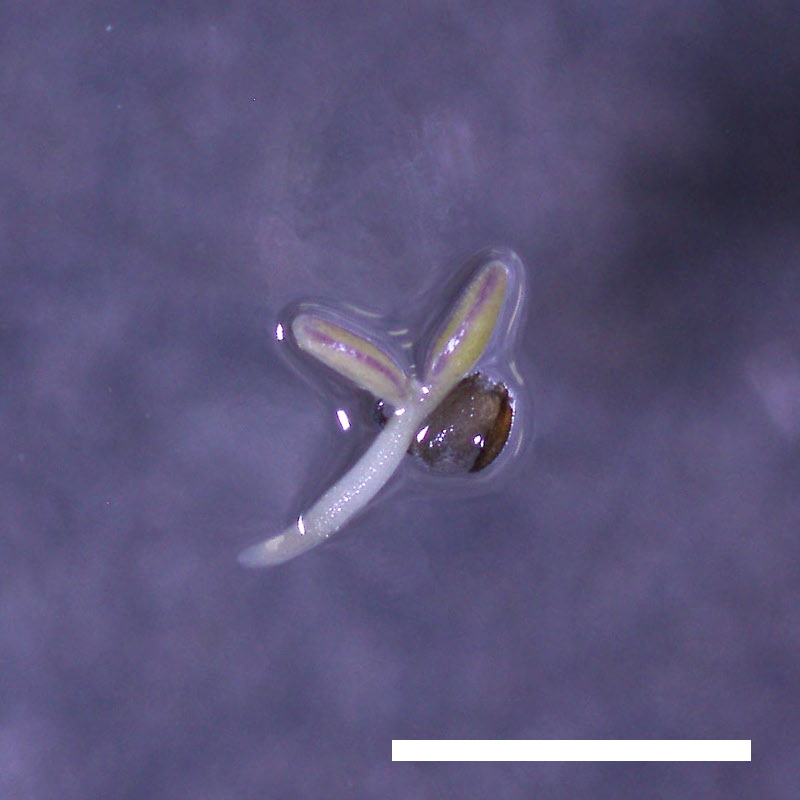
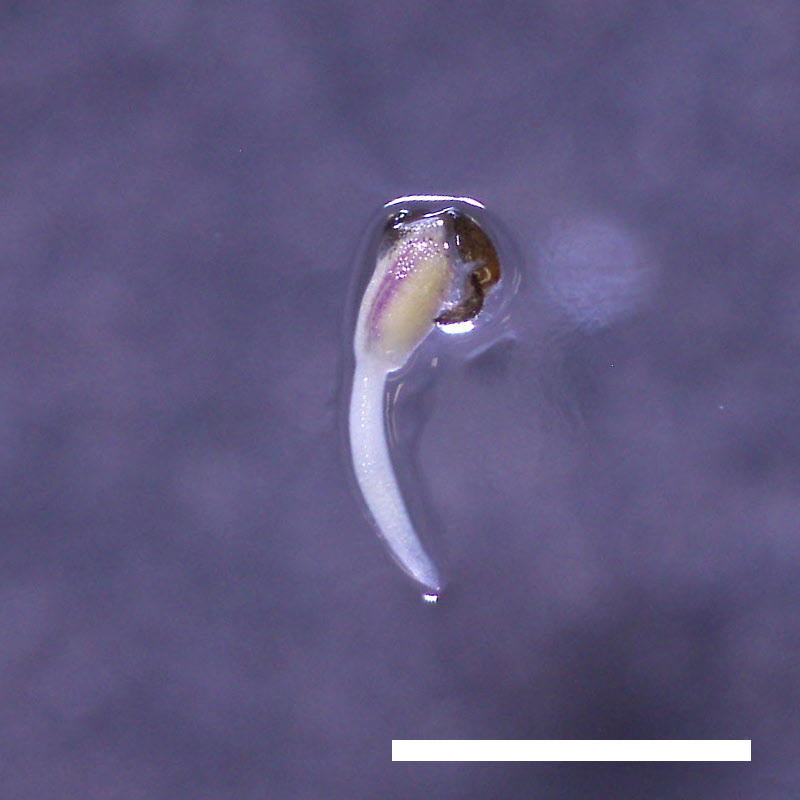
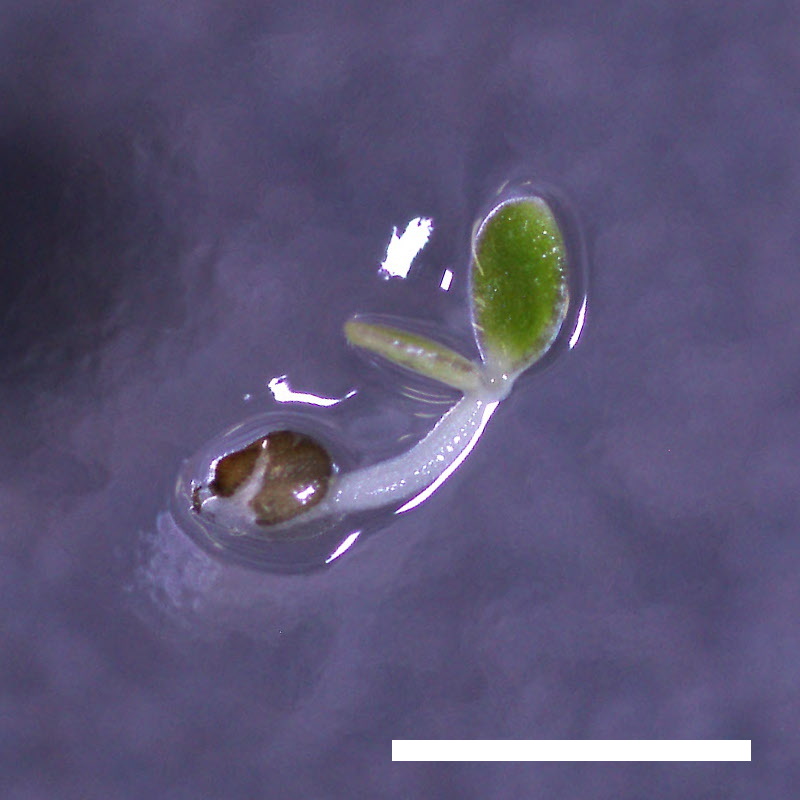
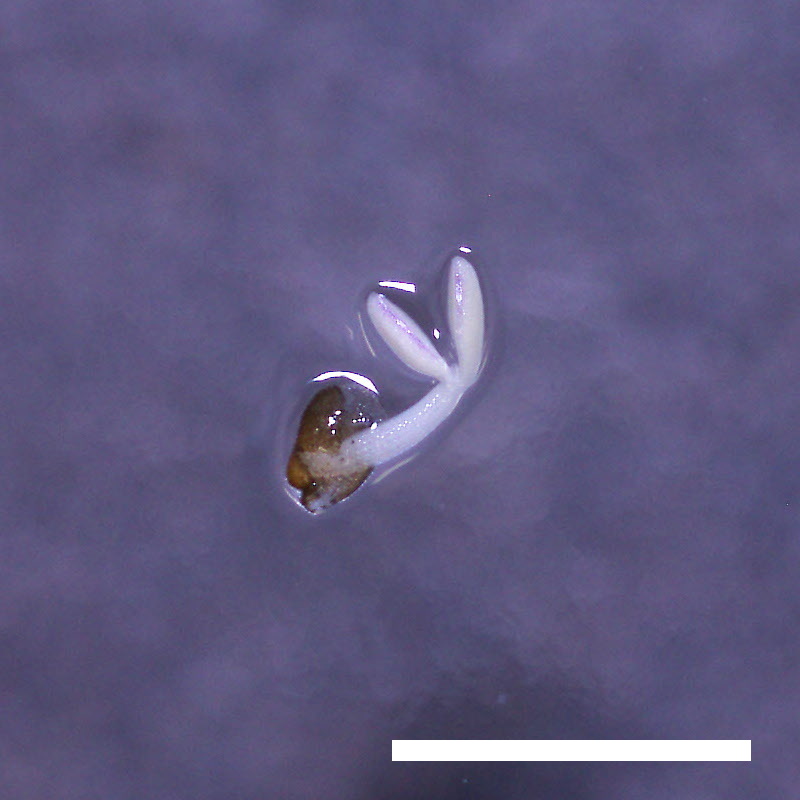
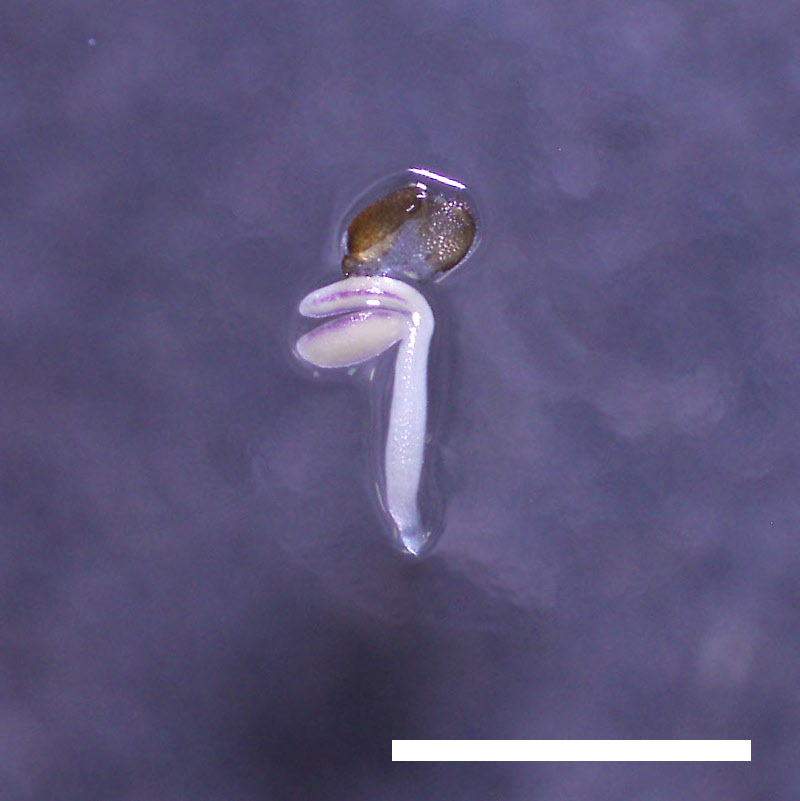
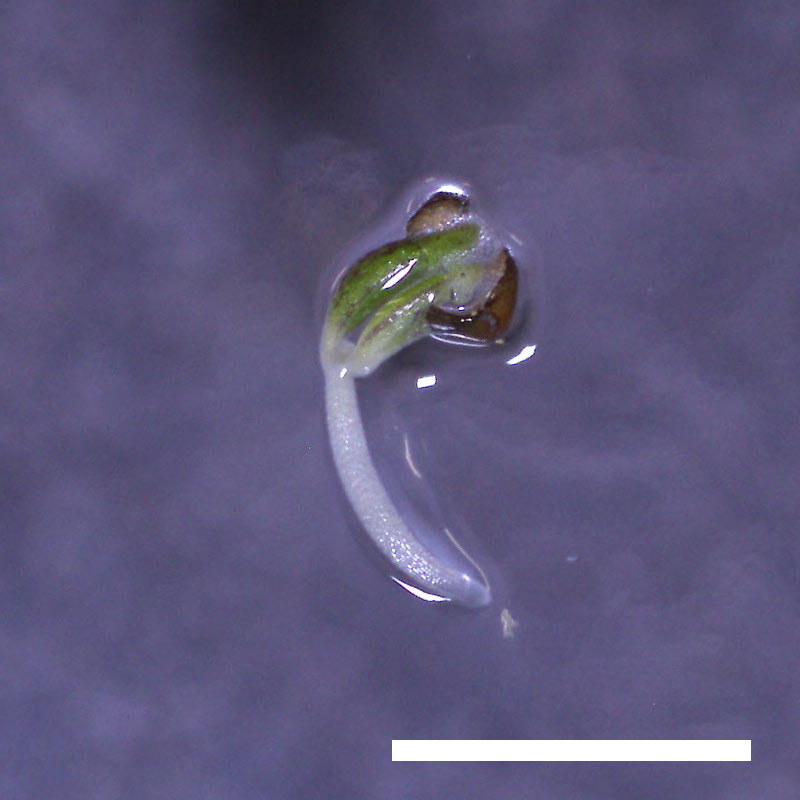
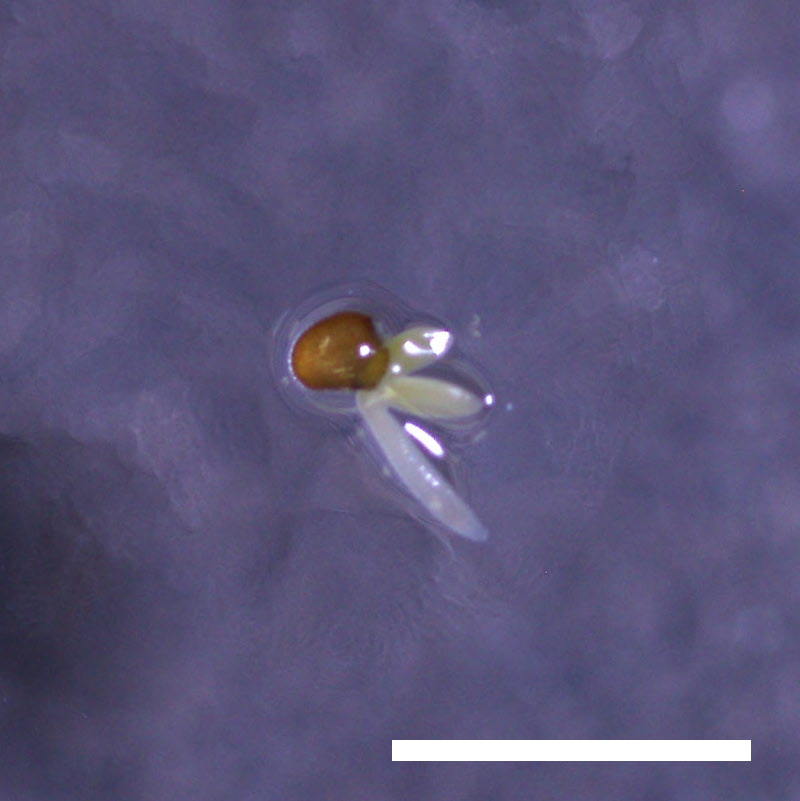
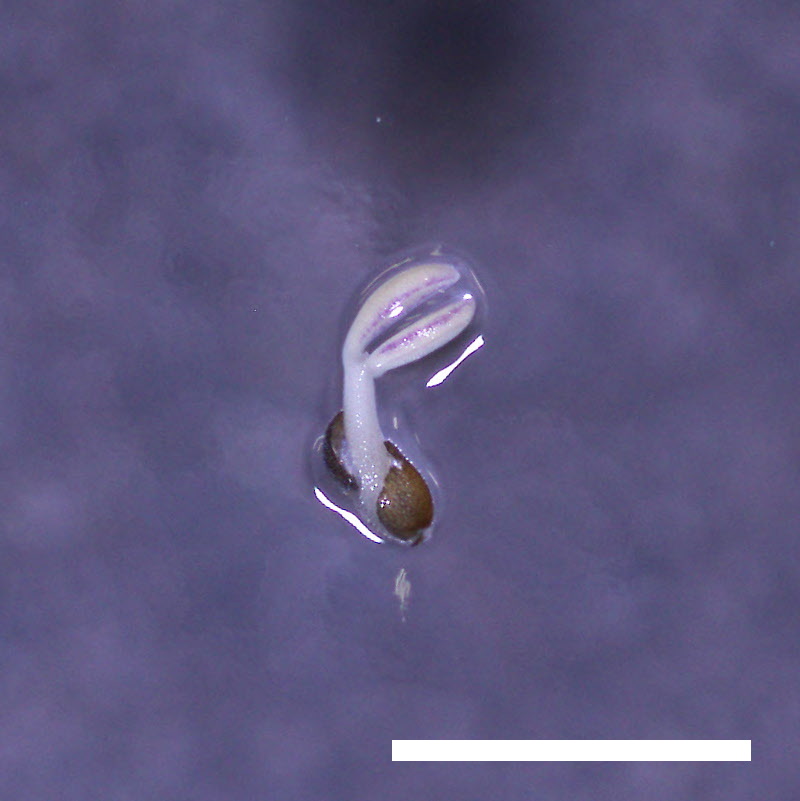
- Opening of the cotyledons was inhibited at 0.5 µM ABA in the drol1 mutants.
- Greening of the cotyledons was inhibited at 1 µM ABA in the drol1 mutants.
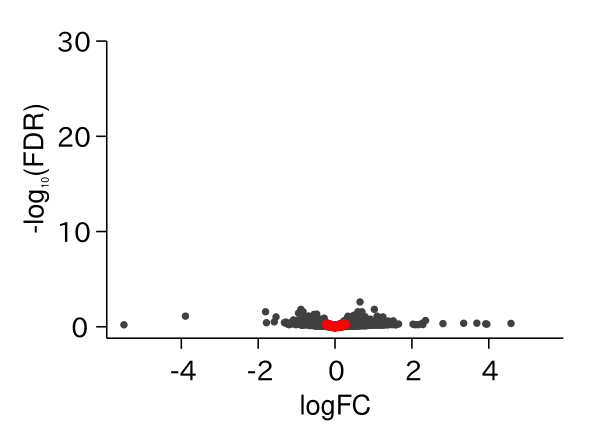
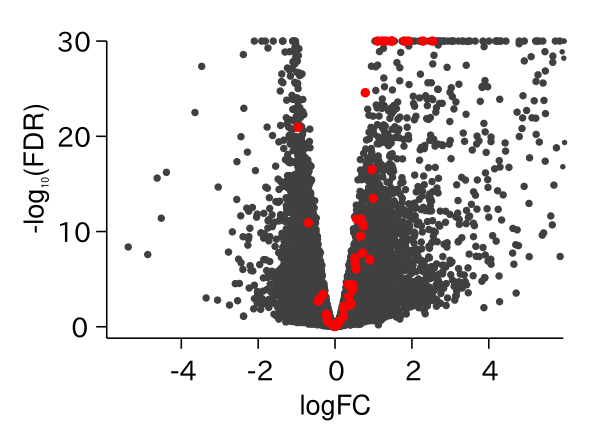
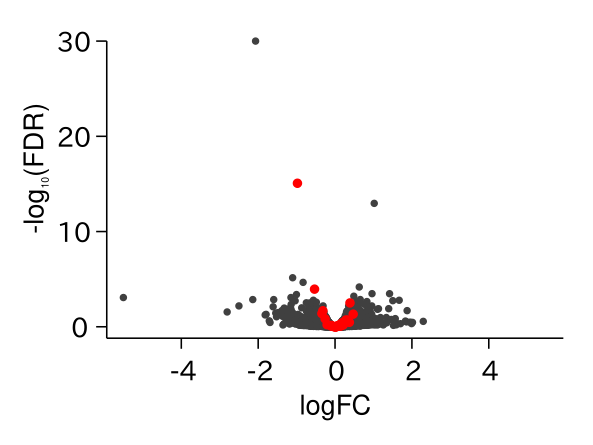
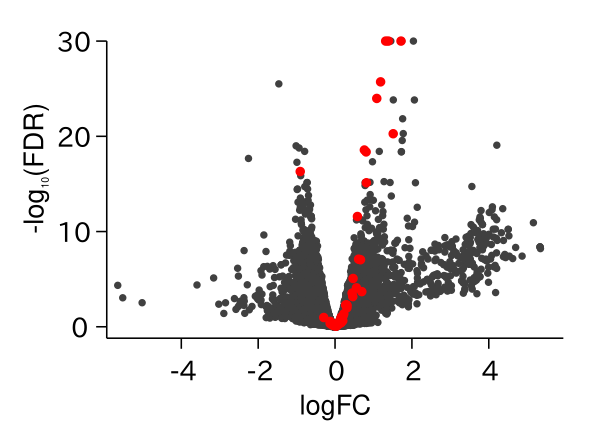
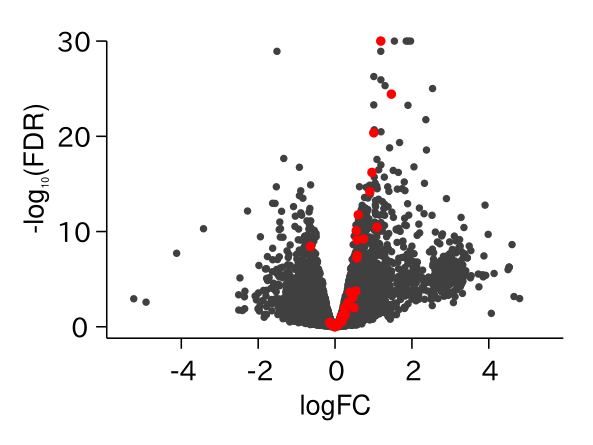
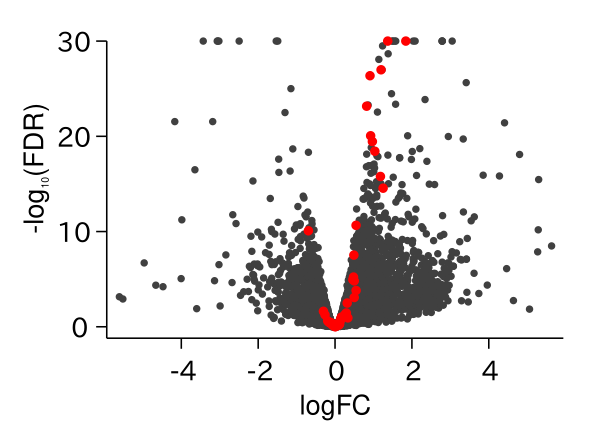
ABA Responsive Genes Were Up-regulated in drol1
Key Question
- どうしてスプライシングの欠損がABAへの応答を引き起こしているのかは不明である
- In drol1, AT–AC introns are specifically retained, and ABA signaling is constitutively activated.
- How does retention of AT–AC introns generate an ABA-like signal?